Abstract
During transcription, the nascent RNA strand can base pair with its template DNA, displacing the non-template strand as ssDNA and forming a structure called an R-loop. R-loops are common across many domains of life and cause DNA damage in certain contexts. In this review, we summarize recent results implicating R-loops as important regulators of cellular processes such as transcription termination, gene regulation, and DNA repair. We also highlight recent work suggesting that R-loops can be problematic to cells as blocks to efficient transcription and replication that trigger the DNA damage response. Finally, we discuss how R-loops may contribute to cancer, neurodegeneration, and inflammatory diseases and compare the available next-generation sequencing-based approaches to map R-loops genome wide.
See more on Pubmed
Read More
Read More
Read More
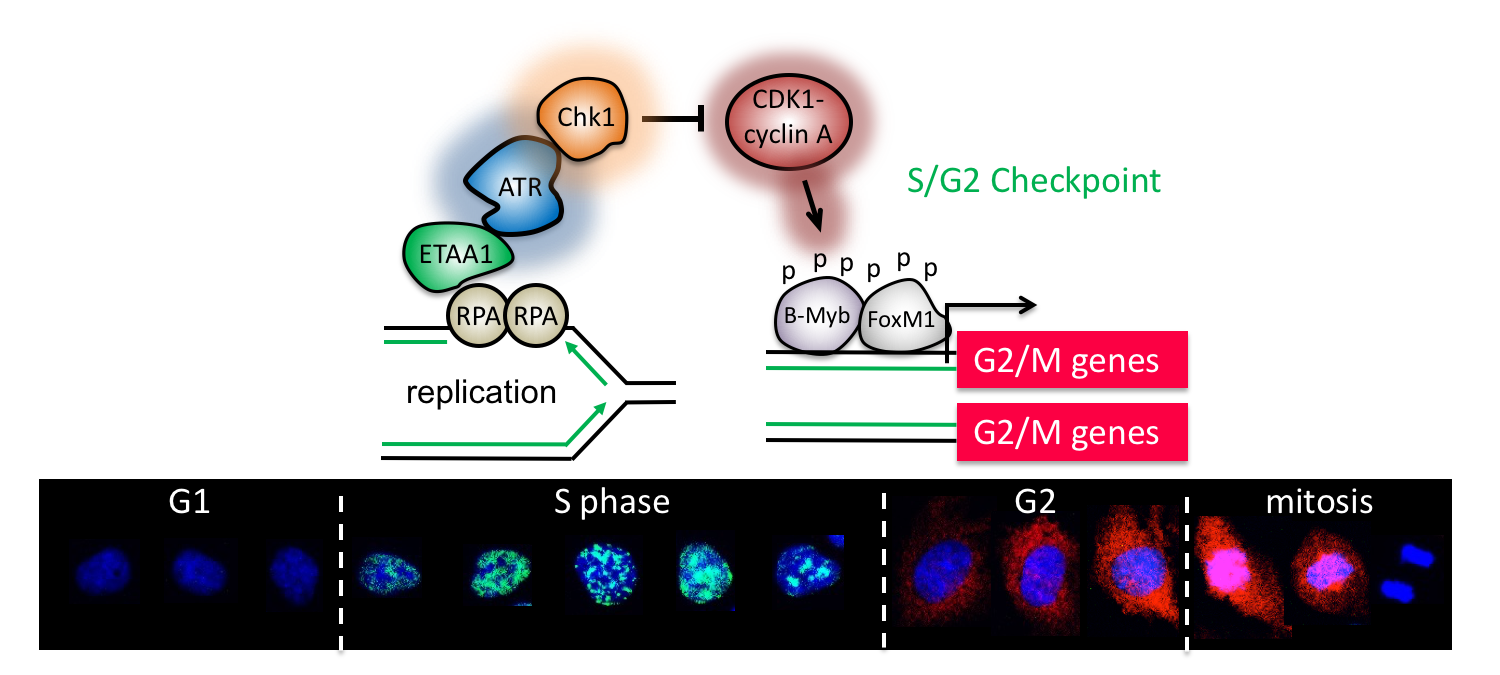 The cell cycle is strictly ordered to ensure faithful genome duplication and chromosome segregation. Control mechanisms establish this order by dictating when a cell transitions from one phase to the next. Much is known about the control of the G1/S, G2/M, and metaphase/anaphase transitions, but thus far, no control mechanism has been identified for the S/G2 transition. Here we show that cells transactivate the mitotic gene network as they exit the S phase through a CDK1 (cyclin-dependent kinase 1)–directed FOXM1 phosphorylation switch. During normal DNA replication, the checkpoint kinase ATR (ataxia-telangiectasia and Rad3-related) is activated by ETAA1 to block this switch until the S phase ends. ATR inhibition prematurely activates FOXM1, deregulating the S/G2 transition and leading to early mitosis, underreplicated DNA, and DNA damage. Thus, ATR couples DNA replication with mitosis and preserves genome integrity by enforcing an S/G2 checkpoint.
The cell cycle is strictly ordered to ensure faithful genome duplication and chromosome segregation. Control mechanisms establish this order by dictating when a cell transitions from one phase to the next. Much is known about the control of the G1/S, G2/M, and metaphase/anaphase transitions, but thus far, no control mechanism has been identified for the S/G2 transition. Here we show that cells transactivate the mitotic gene network as they exit the S phase through a CDK1 (cyclin-dependent kinase 1)–directed FOXM1 phosphorylation switch. During normal DNA replication, the checkpoint kinase ATR (ataxia-telangiectasia and Rad3-related) is activated by ETAA1 to block this switch until the S phase ends. ATR inhibition prematurely activates FOXM1, deregulating the S/G2 transition and leading to early mitosis, underreplicated DNA, and DNA damage. Thus, ATR couples DNA replication with mitosis and preserves genome integrity by enforcing an S/G2 checkpoint.
Link to Article
Read More
Read More
 Read More
Read More
 Read More
Read More
Read More
Read More
Read More
Read More

Dr. Karlene Cimprich was awarded a BRCA Research Collaborative Grant from the V Foundation.
Read More
↑ Back To Top
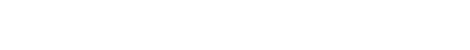
 The cell cycle is strictly ordered to ensure faithful genome duplication and chromosome segregation. Control mechanisms establish this order by dictating when a cell transitions from one phase to the next. Much is known about the control of the G1/S, G2/M, and metaphase/anaphase transitions, but thus far, no control mechanism has been identified for the S/G2 transition. Here we show that cells transactivate the mitotic gene network as they exit the S phase through a CDK1 (cyclin-dependent kinase 1)–directed FOXM1 phosphorylation switch. During normal DNA replication, the checkpoint kinase ATR (ataxia-telangiectasia and Rad3-related) is activated by ETAA1 to block this switch until the S phase ends. ATR inhibition prematurely activates FOXM1, deregulating the S/G2 transition and leading to early mitosis, underreplicated DNA, and DNA damage. Thus, ATR couples DNA replication with mitosis and preserves genome integrity by enforcing an S/G2 checkpoint.
The cell cycle is strictly ordered to ensure faithful genome duplication and chromosome segregation. Control mechanisms establish this order by dictating when a cell transitions from one phase to the next. Much is known about the control of the G1/S, G2/M, and metaphase/anaphase transitions, but thus far, no control mechanism has been identified for the S/G2 transition. Here we show that cells transactivate the mitotic gene network as they exit the S phase through a CDK1 (cyclin-dependent kinase 1)–directed FOXM1 phosphorylation switch. During normal DNA replication, the checkpoint kinase ATR (ataxia-telangiectasia and Rad3-related) is activated by ETAA1 to block this switch until the S phase ends. ATR inhibition prematurely activates FOXM1, deregulating the S/G2 transition and leading to early mitosis, underreplicated DNA, and DNA damage. Thus, ATR couples DNA replication with mitosis and preserves genome integrity by enforcing an S/G2 checkpoint.

